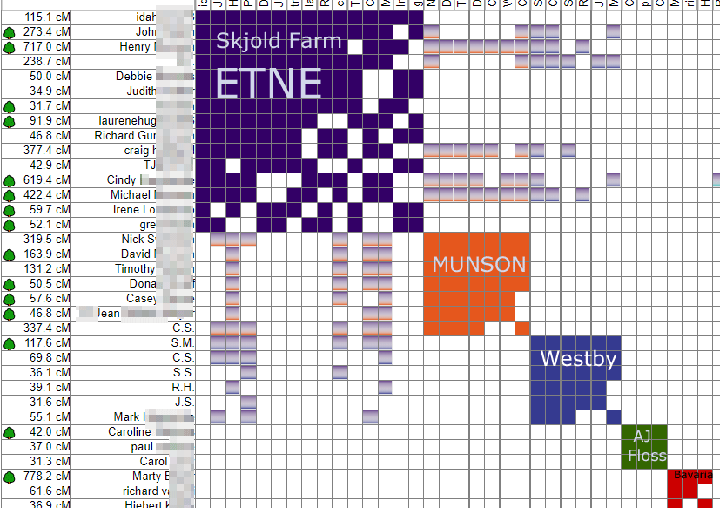
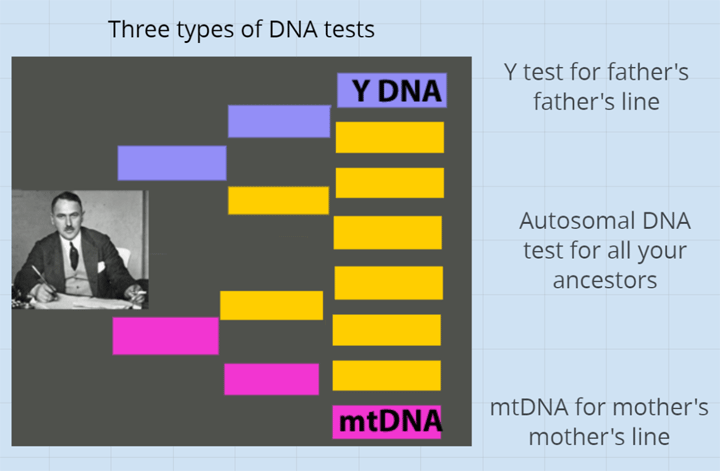
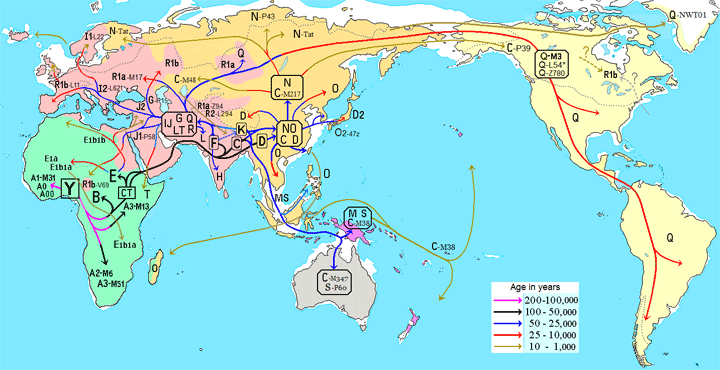
My Dad’s Norwegian great great grandfather, Lars Monsen, was originally from the Bergen area we had been told. He left his ship to marry a girl from Farsund and then settled in Kristiansand. My cousin Dick and I tried to find records about him, but this name was quite common in Hordaland (think Tom Smith). There were 10 candidates for our Lars in the Bergen area, so I used Y DNA testing to figure out which one he was. This is discussed in several previous blog posts (click here and here)

Kristiansand Boat Basin 2015
Sigmund, a friend in Norway, found a paternal line descendant of Ole Monsen Åstveit, my suspected 5rh grandfather. Then Sigmund sent him a Y 37 marker test from Family Tree DNA. This cousin, Einar, and my father subsequently matched at 3 steps, meaning there were three STR markers that were different. Since the common ancestors lived in the 1700s this seemed reasonable, especially after I looked at each of the markers and found them to be faster mutators (click here for that article). On the other hand, my second cousin, also in the R1b haplogroup, matches a 5th cousin of his, descended from an ancestor in the 1600s, at zero steps. Basically any match in the 0-3 range is usually recent enough for the genealogies to line up.
For Christmas this past year, I gave myself the gift of upgrading Einar’s Y STR test to the BigY700 at Family Tree DNA. I had long since upgraded my Dad and that upgrade included more STR markers. For more about the different Y tests try my article Why Y? (irresistible title!) which has links to more resources as well. This upgrade also took Einar’s STR test from 37 markers to 111. Imagine my surprise when I found that at 111 markers they still had only a 3 step difference!
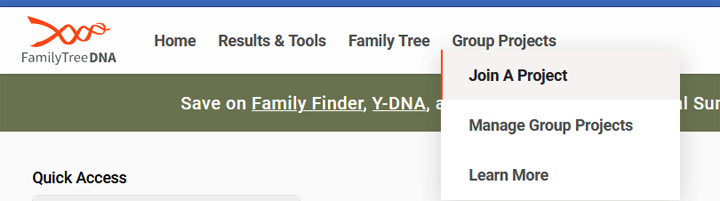
However the purpose of the BigY700 is to look at SNPs rather than the STRs. The SNPs will tell you more about your deeper paternal line ancestry. It is also a good way to confirm a STR match. Another reason to do the big Y is that when you have an active project administrator, you will be able to see the bigger patterns and branches of your section of the Y tree. There are projects for specific surnames as well as for regions and haplogroups and subsets of haplogroups. You can join projects at Family Tree DNA from your dashboard by clicking Group Projects then Join a Project, as shown below.
Continue reading