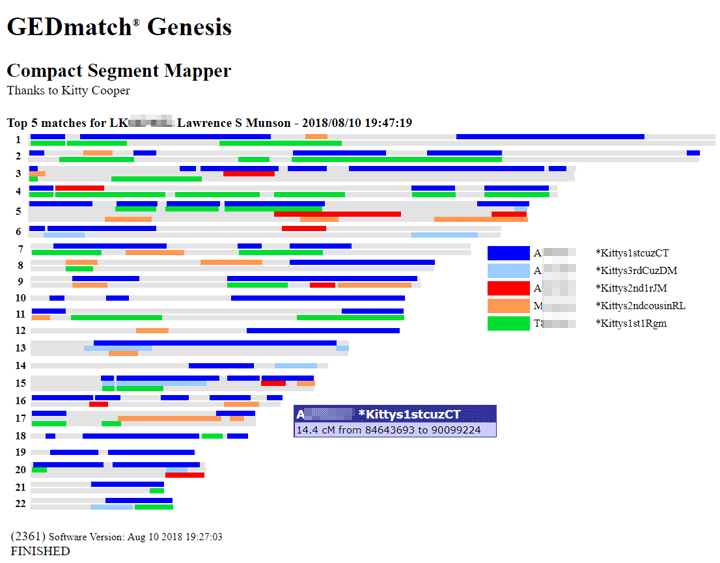
Today GEDmatch launched a version of my segment mapper which is integrated with their database. It can be used as a way to easily visualize multiple matches on GENESIS. This first version shows how everyone compares to the first person on the list, like the current 2D chromosome browser but more compact. Also each color is a person.
Above is the screenshot of my Munson cousins which even shows one of the boxes that appears when the cursor is placed on a segment. Click on this image to see a copy of the html version (edited for privacy) where you can put your cursor on any segment to see the details. (N.B. when you put your cursor on a segment and the information pops up via javascript that is called a mouse over in geek speak. )
One way to get a display like this at Genesis is to check the boxes (tier 1 only) next to your best matches in the one-to-many (up to 40 of them) and then click the button “Visualization” at the top of the page. Another way to get there is to use the Tier 1 tool “MultiKit Analysis” and select the kits you want to look at. Remember the first kit listed is the one that all the others will be compared to.
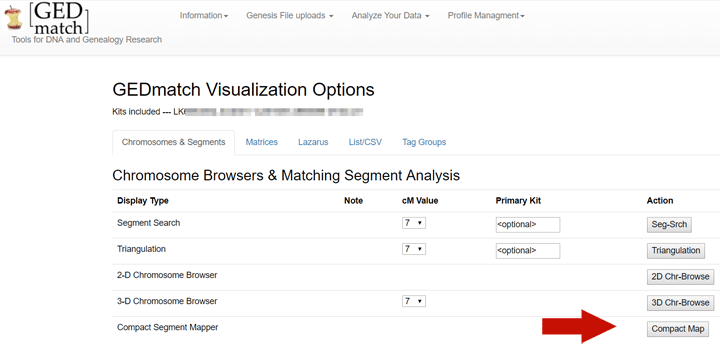
Once you are on the visualization page, at the bottom of the Chromosomes & Segments display there is a button called “Compact Map” which will make a chromosome map like the one above. I put a red arrow next to it in the image below.
If you have not yet played with Genesis, which will ultimately be the main GEDmatch site, here is a reason to try it. Your user id from GEDmatch will work on Genesis. Many kits are already migrated over. If yours is not yet moved over, you could just go aheas and upload your raw DNA file there. As you may already know, new 23andme kits and LivingDNA kits can only be uploaded to Genesis and not the old GEDmatch because they use the new chip.
I confess, I offered to sell my segment mapper code to them as I thought it would vastly improve their display functionality. I also agreed to support it and occasionally do additional work for them. How many of you knew I was a programmer as well as a blogger? And yes I have been doing alot of coding this week which is why I have not been blogging!
My next idea is to include everyone’s match to everyone else with one color per person and then indicate who they match where in those mouse over boxes. This would be a multi-way version of this chromosome browser, so a compact way to see your family triangulations. It is almost ready.
By the way another wonderful feature at Genesis is that you can select a group of kits to use for triangulation in the Tier 1 multi kit analyis. The way that works is when two of those kits have a three way match with the first kit, the match is displayed.
I am really excited about my segment mapper’s new home at Genesis. Give it a try and let me know what you think!

Thanks Kitty, this is an excellent addition to GEDmatch Genesis! I look forward to the further additions to this feature.
Chuck
I don’t have any boxes by my matches to click on.
The checkbox is tiny, right after the kit number on your one to many. Also the tier 1 one to many has it more prominently at the beginning.
I don’t have any boxes to the right of the kit number. I’ve put the mouse there in case they’re too tiny to be able to see with the naked eye, and there’s nothing there . . .
Darlene –
I just double checked by using a non Tier 1 account and I see that the box does not show in that account. Apologies, this is a tier 1 feature
When you inspect with chrome and firefox dev tools there’s no checkbox code or visualization button. Thanks for creating this great tool. I look forward to using it.
As I said above ed, the boxes are apparently tier 1 only. I corrected the post
Someone else reported getting to this from the kits that match 2 kits which has check boxes
Oops, I just saw your reply to Darlene about the checkbox.
Great tool, Kitty. FYI, the details pop-up worked on my iPad, too, with a light touch in place of the ‘mouse-over.’
Cheers,
Kathleen (the other K Cooper)
dear friend other K –
Great to hear that, thanks!! I am thrilled myself, they did a nice job with it!
Thank you for this new tool. If you have done Visual Phasing, you can use both tools together. I compared my first cousins, my paternal great uncle, and my paternal aunt to myself. since i know which chromosomes i inherited from my paternal grandparents, I can see where my cousins, aunt, and paternal great uncle match on which grandparent lines on my chromosomes. i inherited all of chromosome 15 from my paternal grandmother, therefore my paternal great uncle or his daughter do not match me on there (they are from my paternal grandfather’s side). I also use my Triangulation list to verify the matches on the chromosomes. Looking forward to the update for the triangulation feature.
oh, and the cousins i was comparing myself to were also from my paternal side.
I haven’t been on GEDMATCH’s GENESIS in a while, but your post inspired me to try your segment mapper tool. Love this. It’s extremely user-friendly (thank you)!
Extra added bonus is that your name popped up as a match! I did the proverbial double-take when I saw your name. Wow! GENESIS says we’re distant cousin range (sharing 15.1 centimorgans), but I still thought it was very cool.
Thanks Kitty!
Kitty,
It’s wonderful to see your segment mapper being made available to a wider audience. It’s a great tool!
On a programming note, do you know why the starting and ending locations of the segments seldom are displayed with commas separating the thousands? For us with astigmatism, these long numerical strings are a challenge to read without the commas.
Jim,I agree and the beta of the new version rounds to the nearest million so for example 17M to 38M
Thanks, Kitty!
You are a genius! Wish I was a cousin to you too (-:
Wow! GEDmatch is such an awesome tool for genetic genealogisits. As someone who is working on their family heritage with a limited budget, I can say that GEDmatch is outstanding value for the price. To start with, the free tools are an invaluable aid in solving puzzles. The very reasonable price of a Tier 1 membership combines well with non-automatic renewal for a one-month membership. I can plan my investigations for specific months when I have more time, or I can plan my research ahead of paying to make the most efficient use of a one-month purchase, and then buy another one-month membership for queries a little further down the road. I am thrilled to have such another fine tool added to the site.
Pingback: This week’s crème de la crème — August 18, 2018 | Genealogy à la carte
I have a question regarding GEDMATCH info. I am a female and share an X-chromosome with another female. The data clearly indicates a solid blue line determining that we are half sisters (same father). Is it possible that we share the x-chromosome because our fathers are Brothers making us cousins instead of half sisters?
Sherri –
It is possible but extremely unlikely because both brothers would have had to get exactly the same X from their mother which hardly ever happens since she recombines her two X chromosomes when she passes them on.
Look at the total cMs and look at the sizes of the largest segments. If your fathers were brothers you will share much less DNA. See This calculator:
https://dnapainter.com/tools/sharedcmv4
Paternal half siblings will also share several segments >100 cMs see this post:
https://blog.kittycooper.com/2017/09/the-25-relationship-a-first-look-at-the-data/
Hi–It is not clear, to me, if GEDmatch is totally free?
On the web site I looked at there was no reference to Tier 1-
What am I missing? Thanks
jim
Jim –
Gedmatch is totally free but there are extra features that come with a Tier 1 membership. Once you are logged in, scroll to the bottom of any page ad see the button to subscribe
Hi! New to the DNA part of genealogy – have been doing genealogy for a long time, the old fashioned way! I have a question regarding the compact segment mapping. I see on your example above, certain chromosomes have 2 lines which is where my question comes in. I realize that with multiple matches on one chromosome that they will “stack” (for ease, looking at your chromomsome 1) there is a second line…question is is this determining maternal/paternal lines? And if so, is paternal always first? Hope this makes sense! Thanks in advance!
Sorry Debrah, this does not determine maternal versus paternal. The current technology cannot tell them a-part unless you have a parent tested. These segments are just a mtch to the base person (you?) and not necessarily to each other
Thank you!
I love the compact segment mapper, thanks so much for making it available at Genesis. Still unclear on when it chooses to display two separate bars for the chromosome vs 1, though. I have seen it split segments across separate bars that I know fully triangulate – all from close-ish family, no possibility of separate homologs doing independent matching, big-ish segments, same endpoints! Maybe just a subtle SNP density issue or something? Anyway, great job!
Hi,
Disappointed in the new version of GEDmatch as some of the functions previously available have been moved into the Tier 1 group as part of the ‘visualisation’ feature or have not been carried across from the old. In particular the ability to compare multiple people.
While the Tier 1 facilities are nice and I have used them from time to time, it becomes expensive as the USD-AUD exchange rate is unfavourable to buying US products. Shame
Patrick
Patrick –
It is not free for them to run the site so paying $10 USD for a month’s tier 1 access seems very reasonable to me. Cloud storage adds up.
The multi kit analysis tools are tier 1 since mainly the power users want them and they are willing to subscribe. If this is a hardship for you, just do one month as needed or send me why from my contact form and I will consider buying you a month.
Kitty
For some reason, this site wouldn’t take my VISA months/yrs. ago and probably hasn’t changed I don’t want to have to send a check off to someone but I guess I will — and wait. annoying, but I’ll still to see who’s on my list of matches from the free version. I don’t understand much of the rest of it all anyway, so probably a waste of money for me.
When using this tool for visualizing MKA with the GEDmatch Ancestry projects, what is the significance of separate line for a chromosome vs kits all in the same multi-kit, single, taller gray bar? It does not appear to be maternal/paternal in my usage, but possibly multiple kits that are the same in different single height lines.
Steven –
Separate lines are used for clarity when they would otherwise overlap each other on one line. Sometimes those lines are separated by a little white space (wne 2 of them) else it looks like one tall bar.
No significance.