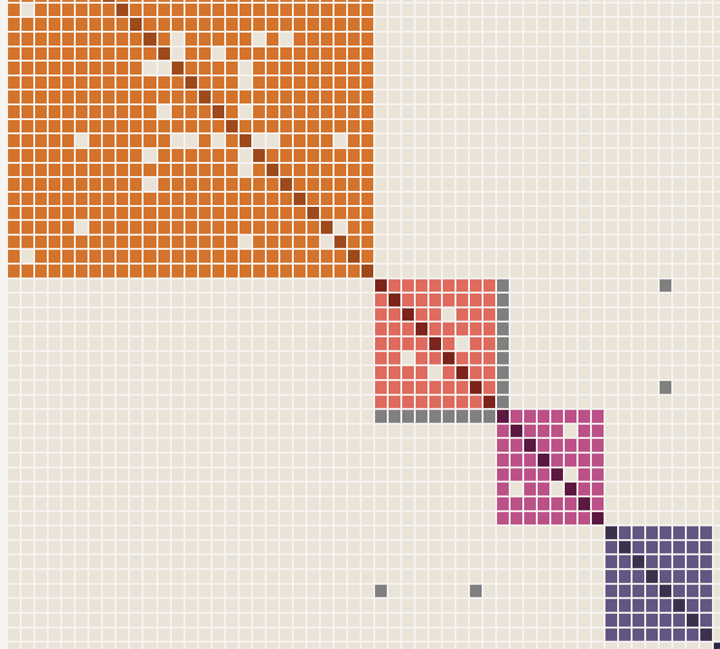Randi contacted me for help finding her unknown biological father. I advised her to test at Ancestry and get back in touch when her results were ready.
When they came in, it took only three hours to find Randi’s unknown biological dad! She had two second cousin level matches at Ancestry with good trees who did not match each other. That meant that the search would likely be easy, since all I had to do was find where those two trees intersected. Here is how I did that, step by step.
First I created a private searchable tree at Ancestry to use for this case. I started it with Randi and her mom. planning to make floating branches for related people by copying the relevant lines over from their trees.
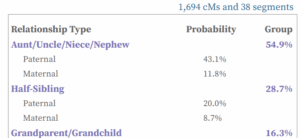
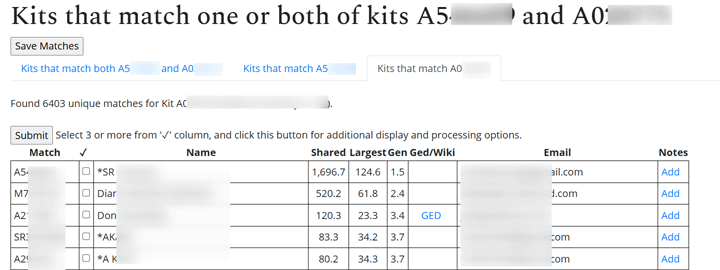
The best DNA match on Randi’s unknown father’s side was Brad at 322 cM. So using ProTools, I sorted the matches Randi and he shared by his closest matches, as shown in the image below.
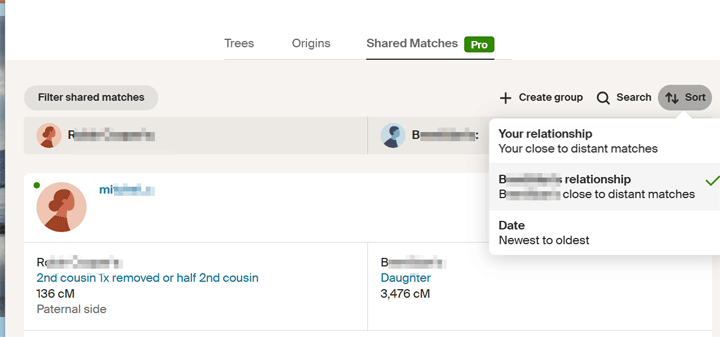
Clicking on the sort button brings up a box where you can select to sort by the match’s relationship
The idea was to find the common ancestor among those matches. This would be the line that Randi is related on. To do that, I should have looked at the best one with a tree, excluding close family, but I saw that there was one a bit further down the list, Bob, who had an unusual surname, call it Roper, that was the same as one of Brad’s great grandmothers. So I built her tree back up a few generations and down again. I then copied Bob’s paternal ancestors over, looking for an intersection. I did not find one, so I moved on. Later Brad told me that the error was two women with the same name and birth year incorrectly in various trees, including his.
So I went back to the common match list and found the best match to Brad with a tree (Peggy). One of her grandmothers shared a surname, call it Whistler, with the husband of the Roper great grandmother. So I built the Whistler tree. Quickly found a common ancestor for Peggy and Brad with an unusual first name born in 1830. Built the tree of all his descendants. Somewhere in that tree will be our man.
Time to look at the other possible second cousin match, Jim, at 268 cM. The plan was to repeat the process of finding the common ancestor to his best match with a tree. However, his surname, call it Wander, had already showed up in the Whistler tree. Having collected all the descendants of Brad’s Whistler great grandparents, I noticed that one of them had married a Wander. Was that Wander in Jim’s tree? Yes, she was his aunt!
That Whistler-Wander couple, who must be Randi’s ancestors, had two children, a boy and a girl. The boy was the right age and in the right location. Could it really be this easy? Yes. The details of his life fit what was known. Since Randi’s presumed father is Jim’s first cousin, Jim is Randi’s first cousin once removed.