Technology never stands still. The latest change affecting all of us who love using DNA for genealogy is a new chip from Illumina. The past six or so years of autosomal DNA testing have shown that the current chip is great for testers with European ancestry, but does not have enough SNP coverage to figure out the details of the ethnic make up for people from other parts of the world. Many more and different SNPs are tested in this new GSA chip.
All the 23andMe tests done since this past July use that chip, as does Living DNA (highly recommended if you have British ancestry since it does local regional breakdowns). I imagine eventually the others will follow along. The bad news is that there is not that much overlap between this chip and the previous ones, which affects cousin matching.
Debbie Kennet wrote a blog post describing the new chip at https://cruwys.blogspot.com/2017/08/23andme-launch-new-v5-chip-and-revise.html
Because the SNPs are so different the DNA results from these kits cannot be uploaded to GEDmatch, however our friends there have built another site to handle these new kits called GENESIS. They have come up with a whole new algorithm for relative matching that works with lower SNP counts.
At the recent i4GG.org conference (videos coming in February), I gave a presentation on what’s new at GEDmatch, the second part of it went into much detail about GENESIS, starting with this slide:
http://slides.com/kittycooper/gedmatch-10-13-23#/38
The functions available at GEDmatch are being gradually implemented at GENESIS. Most of the key ones are there now. Plus there is some new functionality. One major addition is the showing of the number of SNPs actually overlapping between kits. Very important to know since the overlaps can be as small as 108,000 SNPs or as large as 580.000.
The thing I like best is the new multi kit analysis selection page which lets you add kits easily, including dropdowns of your own kits, to the ones that you already checked at the one-to-many or kits that match two kits. But I miss the tag groups.
The other new feature that is great is the ability to look just at the FIRs (fully identical regions) on the one to one comparisons. Only full siblings and other close doubly related people share long blocks of FIRs. It also gives the total number of cMs in those FIR blocks, which is very helpful for figuring those ¾ relationships. UPDATE 16-Dec-2017: I have received emails asking about the statement they see under a one to one match like “80.8 Pct SNPs are full identical” – these are individual SNPs, normal when you come from similar population groups. This is not the long shared regions of double relatives.
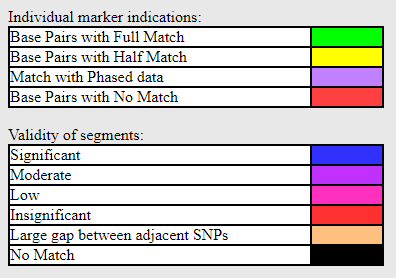 Also new is the addition of more colors for validity on the one-to-one images including a color for too large a gap between SNPs.
Also new is the addition of more colors for validity on the one-to-one images including a color for too large a gap between SNPs.
Missing is the connection to trees from the one-to-many but that will come. The current one-to-many is not the final version I am told.
I recommend you look through the screenshots from my presentation or upload a kit or two and take a look around. In a few months the GEDmatch database will be moved over but why wait?
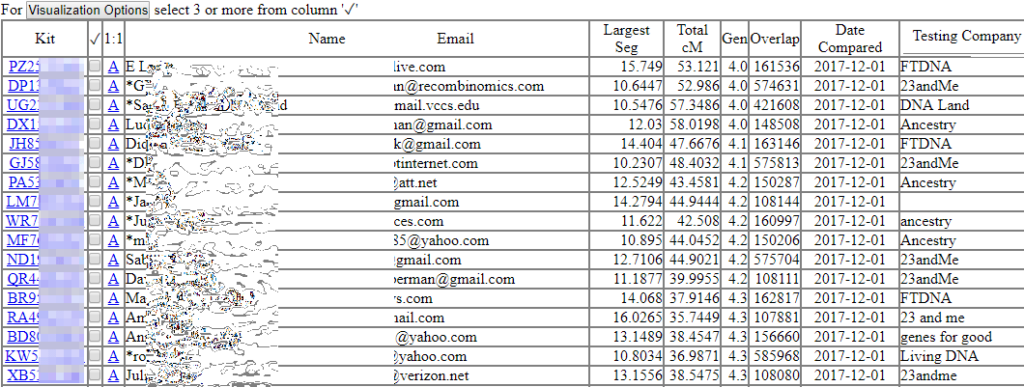
In general, I rested at the two major companies two years ago. Will I need to retest in order to take advantage of these new technologies?
Thank you
You don’t need to, but everything is on sale so you might get a kit that uses the new chip, particularly if you have ancestry from the British Isles try LivingDNA.. They also give your haplogroups
Is the LivingDNA useful if one’s heritage is from the Connemara region of Galway, Ireland? My ethnic breakdowns from Ancestry and FTDNA indicate that I have “British Isles” in addition to Irish/Scottish.
Barbara –
Ask them but yes it might well be. One thing is that when you get your results, you have to click the plus sign next to regional to see the sub regions which many people miss.
Also it gives you your mtDNA haplogroup
Kitty
Thanks for the tip on Living DNA (during a sale no less)! Roughly half my ancestry is British. Very interested to see this breakdown.
Kitty, I’m with you – love those tag groups!
🙂
“Only full siblings and other close doubly related people have these. [FIRs]” — Just to be clear for newcomers – you are addressing very large FIRs.
We all have smaller regions of FIRs with other humans, it’s why false-positives are so common in half-identical matching.
As for Genesis itself – I don’t know how successful they will be able to overcome the low commonality some genotype files will have with others, especially as newer genotyping products come and go and use different SNP sets. When we get down to only say 100k shared SNPs over all 23 chromosomes.
I guess time will tell.
Dan –
Thank you for pointing out that my wording was not clear. I have fixed that, I hope, and also added an explanation for another point of confusion from several readers (see the update in the FIR paragraph).
Another question: I have all of our kits already uploaded to GEDMATCH. Should I re-upload them to GENESIS?
Barbara –
Up to you whether you reupload. Depends whether you want to play with the new features.
Gedmatch says to wait. They will combine the two systems so your old kits will be moved by them. If you do it, your file will be there twice creating a heavier load than necessary.
D Gjervold, I hope you are correct as it is the answer that I have been looking forward to finding. It would only make sense that they do this. Thank you for your posting.
I am concerned. I’m scheduled to give a few talks on gedmatch in early 2018, including one at the Ohio Genealogical Society conference in mid-April. The syllabus for printing is due by mid-January. Now I’m getting that sick feeling that everything I’ve prepared will no longer be valid. Advice?
Kathleen,
If the conference uses an app, you can upload a new version of your syllabus … by mid April I would expect that GEDMatch will have migrated to the full GENESIS functionality … se be prepared? Email gedmatch at gmail for the expected timeline. Meanwhile get familiar with GENESIS and be ready with some of those screenshots!
I have learned much by reading your blog, it has been a huge learning tool. That said, I was on Gedmatch and I have a match to a Kitty Cooper @ 6.8 mrca. If this is your kit, I am honored and a little giddy to be related (and I did double check the match one on one)
Happy new year and thank you for the excellent tutorials.
Beatrice
*Gedmatch Genesis
Both GEDMatch and Genesis don’t allow segment matching at the level of 3-4 cM and with less than 500 SNP. I am attempting to triangulate among known cousins and a potential 8th cousin to establish a common ancestor from early 1700’s. We have a matching segment among three cousins and this potential 8th cousin, but I’d like to determine a probability of the relationship. I hope to recruit more verified cousins to see if we all match on the same segment, and also that at say a 1/100 probability, a random kit will not match. Suggestions? Thank you.
Please don’t use such small segments. They are just not a reliable indicator of relatedness. You can find 3cM matches to almost anyone with North European ancestry.
I do drop down to 5cM and 400 SNPs for KNOWN RELATIVES only but take any small segments with a grain of salt uness they triangulate.
You will only match half your 4th cousins and maybe 15% of your 5ths. See https://isogg.org/wiki/Cousin_statistics
So for deeper ancestry, Y DNA matching is the best tool
Any prediction of when GEDmatch will merge their main database with Genesis? I have a new kit but no clear matches on Genesis, but know that I should be getting a lot of hits on the main database, so am chomping at the bit for when they will be merged. There’s no information at all on the GEDmatch site.
Daniel – I asked them and they hope for soon but cannot commit to a date yet, sorry! I will blog it out and add to the comments here when that happens.
I have no idea what to do. I know I have Native American blood on my father’s side, but it didn’t show up. Was told to upload to Gedmatch. I tried, but not sure if I was successful. I don’t know where or how to look. Technically challenged is an understatement!!
Diana –
See if you can get a technically savvy friend or relative to help you.
The first thing is to create a login id at GEDmatch. When you log in, you can see any kit numbers that you uploaded succcessfully in the left column.
If you tested at 23andme or LivingDNA you need to do the upload at GENESIS. Else GEDmatch.
Here are the instructions for uplaoding an ancestry test:
http://blog.kittycooper.com/2015/09/please-upload-your-ancestry-raw-data-to-a-site-with-a-chromosome-browser/
See if these presentation slides help you navigate GEDmatch:
http://slides.com/kittycooper/gedmatch-10-13#/
I’ll try. Thanks.
I tested with ancestry, uploaded dna to myheritage, ftdna, dnaland, gedmatch, and gedmatch genesis.
My question is, I had a search angel help find my family. I have been in touch with a half-sister, I assume. We figre there are at least 10 sibling in the family, so I thought, to cover all bases, she should test with 23andme. So I bought the kit and had it sent to her. So, to see if we are indeed siblings, will she have to upload her raw dna to gedmatch genesis also?
Lori –
Yes, either that or you test at 23andme
Kitty,
Is there any way to upload Y-DNA to Genesis? I am getting conflicting reports from useres. If not are there any plans to add Y-DNA functionality in the fiuture?
NO, GEDmatch does not do Y matching. If you have tested STRs, then you can upload to http://www.ysearch.org for matches. You can do this automatically if you tested at Family Tree DNA from the bottom of your matches page.
The huge problem I’m seeing with Genesis (unless I’m missing it) is the lack of link to GEDcoms/pedigree charts. Without an ability to find matches this way, at least to start buidling a framework, it’s been useless to my mom who recently tested with 23andMe. My dad test awhile ago and can use the legacy GedMatch – I’ve been able to make many definitate relationship connections which I can use as “standards” when comparing other unknown relationship matches. My mom is going to re-do her testing so she can get a legacy kit and access to the pedigree charts via GEDcoms files. I’m also encouraging folks to put their 5-6 gen pedigree chart and GedMatch Kit # in WikiTree to help in relationship finding as well.
Patience Jan, eventually Genesis and Gedmatch will be combined and the tree information will be back.
I recommend testing at Ancestry for people whose main interest is relative matching. They have by far the largest database and the best automated tree matching.
HI Kitty,
Thank you for the article.
A recent adoption case I am working on tested at 23andME and uploaded to Genesis.
For his highest match of about 100 cM, he has found 3 segments that appear on Genesis that do not appear on 23andME. And has found one segment for that test taker on 23andME that does not appear on Genesis.
I am waiting to hear back about the size of the segments.
Thoughts?
Thank you
Debbie
PS he will be testing at ancestry.
Probably the match was on the older chip at 23andme and the algorithm that 23andme uses to fill in the gaps is different from the one GEDmatch is using. If you email me the two kit numbers and a screen shot from 23andme I can pass it along to the GEDmatch folk
Deatails of 23andME and Genesis mismatch from above question
chromosome 23andme cM genesis gedmatch cM
1 7.2
2 19.36 18.7
4 8.33 9.5
4 7.39 7.9
4 8.69 8.7
5 8.4 10
7 6.29 7.5
8 6.8 7.7
8 9.9
9 7.4
11 7.04
19 9.1
sum 72.3 103.6
I uploaded Ancestry.com DNA to GEDmatch and got a kit number. Does that kit number work on Genesis or do I need to upload my dna to Genesis?
Thank you. Rebecca Duncan
Recently?
All uploads have to be done at GENESIS these days.
Genesis kits start with 2 letters. An ancestry kit uploaded to GEDmatch will start with one letter, an “A”
I recommend uploading to Genesis and then if your migrated kit was already theres mark that old one as “research”
Hi, it seems FTDNA is now taking 23andme v5 and I’ve read comments in a Facebook group suggesting FTDNA is using GSA chip as of 2019 (Jan or Feb).
I imported my mother’s 23andme v5 raw data into FTDNA, here is what I observed after processing completed today:
– 23andme v5 imported into FTDNA, in April-2019, had a total of 752 matches, of which 406 were not in common with same person’s Family Finder results.
– This person’s FF order completed in Dec-2015 and as of April-2019 it has 401 matches, of which 56 are not in common with the recently imported 23andme v5.
Interesting results, need to dig into the information to try and understand what it means.
Thanks Mario for that information. One of the problems with ftDNA is that they report many very small matching segments and ususally over half of those are false, see http://blog.kittycooper.com/2014/10/when-is-a-dna-segment-match-a-real-match-ibd-or-ibs-or-ibc/ for more about false matches.
Still if you discover anything of interest in your digging, please do report back!
Pingback: Friday’s Family History Finds | Empty Branches on the Family Tree