Those of us who are tested at 23andme and have also done the Y STR test at family tree DNA may wonder when some family tree DNA project manager says “Test SNP so and so” whether that SNP is already tested by 23andme. This post explains how to figure that out. If I have already lost you, then this post may just be too technical or else not your cup of tea. To better understand Y testing read this Y lesson by Kelly Wheaton.
For a good explanation of what a STR versus a SNP is, read Roberta Estes’ post – http://dna-explained.com/2014/02/10/strs-vs-snps-multiple-dna-personalities
So to figure out which SNPs my Dad has already tested, I first created the L11 subset image below of the R1b Y haplogroup SNPs from the beautiful diagram created for R1b by Mike Walsh because I need visuals:
Back to the original question. My Dad is an R1b etc and 23andme uses a four year old haplogroup designator rather than the current ISOGG R haplogroup listing. A visitor to this blog suggested that we test DF100 because that is an interesting subclade we may belong to since we have these SNPs according to 23andme: L11/PF6539/S127, L52/PF6541, P310/PF6546/S129, P311/PF6545/S128.
The diagram shows that the possible downstream SNPs for Dad are U106, DF100, and P312. So how to find out if they are tested at 23andme? Since the haplogroup at 23andme shows L52 as the last SNP can I assume the others are tested?
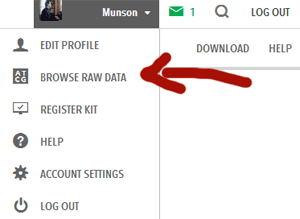 No of course one cannot make such an assumption. New SNPs are found every day and put on the chart.
No of course one cannot make such an assumption. New SNPs are found every day and put on the chart.
There is a way to find the value of specific SNPs at 23andme by looking at the actual DNA data. On the very top left of the screen where your name and picture are shown, you can get a special menu as shown to the left here, by clicking on your name, picture or the down arrow. Click on BROWSE RAW DATA in that drop down menu.
The next problem will be turning a SNP name like U106 into a position on the Y or an “rs” number. To do that, search this ISOGG page of SNPs – http://www.isogg.org/tree/ISOGG_YDNA_SNP_Index.html for the name. For our example, U106, we see this line which tells us that C is the ancestral value and T is the new mutated value and that the “rs” number is rs16981293.
| M405 | R1b1a2a1a1 | S21; U106 | rs16981293 | 8796078 | C->T |
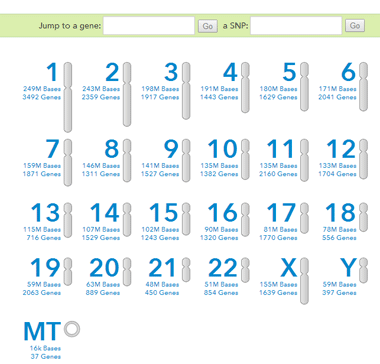
Now we can use that “rs” number on the raw data page. Put rs16981293 into the second box where it asks for “a SNP:” and click go
Some SNPs may not have an “rs” designator. In that case you can use this url and replace the 8796078 for U106 with the one for the location of the SNP you want: https://www.23andme.com/you/explorer/chr/?chr=Y&pos_start=8796078
Let’s say you use the “rs” number, then you will an output page that shows whether you have an A,T,C, G, no call or deletion there.
| Gene | Position | SNP | Versions | Munson’s Genotype | |
|---|---|---|---|---|---|
 |
intergenic | 8796078 | rs16981293 | C or T |
C
|
So as you can see U106 has a C and the ISOGG chart with all the SNPs indicates that this is the ancestral value. So not that branch. DF100 is not tested by 23andme nor is P312. For P312 we need to look at the three downstream SNPs: DF27, U152 and L21. U152/S28 is rs1236440; that SNP is a C which is ancestral, not that branch then. L21/S145 is rs11799226; that SNP is a C which is ancestral, so not that branch then either. DF27 does not have an rs number and using our handy URL for its location https://www.23andme.com/you/explorer/chr/?chr=Y&pos_start=21380200 we find nothing there. So downstream we go again.
Please note that our test results are for the V3 chip, not the newer V4 chip. We have concluded that U106 is tested but not P312 or DF100 at 23andme. However those P312s that are either L21 or U152 do have those downstream SNPs tested. So only the DF27 subset of P312 needs further examining in our case.
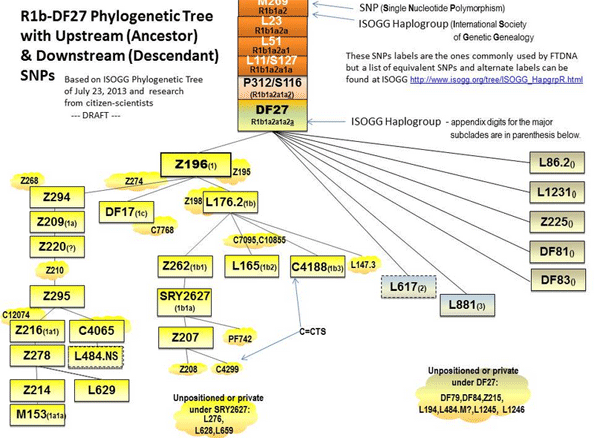
Here is the Mike Walsh chart for DF27 :
The major downstream SNP is Z196 and we see no “rs” number but locations which 23andme did not test
| Z196 | 21033704..21033705 | AT->del |
Z294 is not mentioned on the ISSOGG chart. Z274 is not tested at 23andme. L86.2 – rs9785801 – is not tested. L1231 – 4873643 – not tested.
I will continue working through our results, but I am posting this incomplete since others just want to know how to do this …
not tested: Z198, DF17, Z225, Df81, Df83, L617, L881,
[Ed note: Sometime after this was written, I found an automated tool to do this analysis for me, see that newer post on Y tools ]
Hi Kitty. There’s is an a on the end of my name:) Thanks. Robert+a
So sorry about that Roberta, all fixed! Great blog post by the way, I really enjoyed it.
R-P310 here U106x and waiting for results on P312 and DF100. Good stuff you have posted here.
And the test is in, we are not DF100
Now to figure what SNPs to test for the P312, I guess we can test DF27
I was listed as the same Y Haplo as you. When I transferred my Geno 2.0 to FTDNA it came up as Z198. So P312+>DF27+>Z196+>Z198+ . This is apparently parallel to L176.2 on ISOGG 2014
Thanks Stuart
So now I guess we need to either test DF27 or … Jump downstream? Waiting for advice from R1b experts
The best thing to do is order a comprehensive test such as Geno 2.0, Chromo2, BigY, or FullGenomes. That way multiple SNP markers are tested and the location on the current ISOGG tree can be determined. It’s more expensive to go the one at a time route.
A thread on 23andme announced an automated tool to get all your 23andme Y SNPs – see my blog post on that http://blog.kittycooper.com/2014/04/a-web-site-with-tools-for-y-and-other-dna-analysis/
You are right for most folk Armando but since we expect to be negative for P312 (now ordered), we will be done. If we are positive then we should have gone the other route and I will try and talk my brother into getting one of those!
Why do you expect him to be negative for P312?
Armando, perhaps wishful, we belong to a Facebook group for south Baltic Sea R1b DNA where like us, everyone has L11, P310, P311 and is negative for P312 and U106 so my thinking was that the P310, P311 was predictive for no P312 but I will know in a few weeks.
Also there is our ancestral location of Southern Norway, near Bergen, which may indicate the south Baltic group unless it is Scottish (lots of back and forth with Scotland way back when and we have a few Scottish autosomal DNA matches)
Update: Dad is positive for P312, so now to convince a male Munson to do the NatGEO 2.0 I think …
I’m not surprised he turned out to be P312. That is the largest group in Western Europe under M269. That is especially true for U106- Western European M269. You could always do one SNP at a time. You could take a chance and skip DF27 and test Z195. If positive then test L176.2. If negative for Z195 then test DF27. Then go from there. No idea when the sample will run out though.
The other option is to wait until the end of the year to see if a new Deep Clade test will be offered by FTDNA .
I recently got my ancestry.com DNA results recently and I’m wondering how I find out which haplogroup etc from the raw data..
Kind regards
Ger
Hi Ger-
For the Y I think it is doable … try this post for how
http://www.geneticgenealogist.net/2016/01/how-to-get-ydna-haplogroup-from.html
no automatic way for mtDNA that I know of although Ann Turner can give a good guesstimate …
Hello Kitty I got some information earlier tonight saying Thomas Blackstone b.1615 England to 1645 York Virginia, it says I share Y Chromosome test, haplogroups R-P310, my 23andme Paternal R1b1b2a1a Maternal J1c2 Gedmatch kit # M416339, I have been with a new group trying to understand this Chromosome Mapping and just didn’t know if you could send me some help this way it seems like very nice group of people and I just would like to learn more about this , DNA I got on WikiTree I line up with a lot of Royalty Descendent’s and not so much Native American and it seems if they was a War going on the entire family got involved, Thanks, and have a Blessed day hope to here back
Donnie –
Y DNA is a different test from the autosomal which is what you put on GEDmatch. The Y is your surname line, passes via males only, and is very accurate when enough markers are tested. 23andme tests the Y haplogroup not the markers, as part of its autosomal test.
DNA is fascinating stuff but takes a while to get the hang of it all. I suggest you work through Kelly Wheaton’s lessons which are linked to from my Basics article at http://blog.kittycooper.com/dna-testing/dna-basics/
Kitty
Hello Kitty, I am trying to understand which result is the most accurate between the two different results I got. I tested Y-DNA67 markers and the specific SNP for R-M512 at FamilytreeDna with the result being Confirmed Haplogroup of R-M512. But on 23andme I got R-M417. I am aware that each company may use a different testing method (SNP vs STR) to arrive at the result. So bottom line which is the most accurate? I note that I see R-417 is downstream of R-M412.
Would be great to know since I’m trying to verify the patrilineal lineage on my line which appears to include an NPE. Thanks so much – love your blog and insights!
Jason