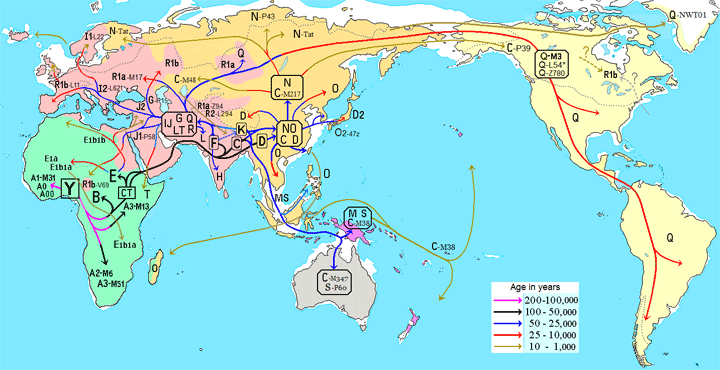
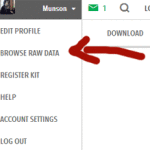
Haplogroups fascinate me because they reveal our deepest ancestry. A haplogroup is a way of assigning a portion of your DNA to a category based on areas of very slowly changing markers. There are two types of DNA that can be assigned haplogroups, because they do not recombine therefore change only slowly via mutations. These are the Y chromosome and the DNA of your mitochondria (mtDNA), which are separate organisms in every cell that provide us with energy and are passed along via a mother’s egg. The groupings for their haplogroups look like family trees when charted, for example the one shown below from Eupedia. That is because each mutation creates a new branch. There are haplogroups assigned for both the all female line (mtDNA) and the all male line (Y). Click here for Eupedia’s wonderful descriptions of all the haplogroups found in Europe.
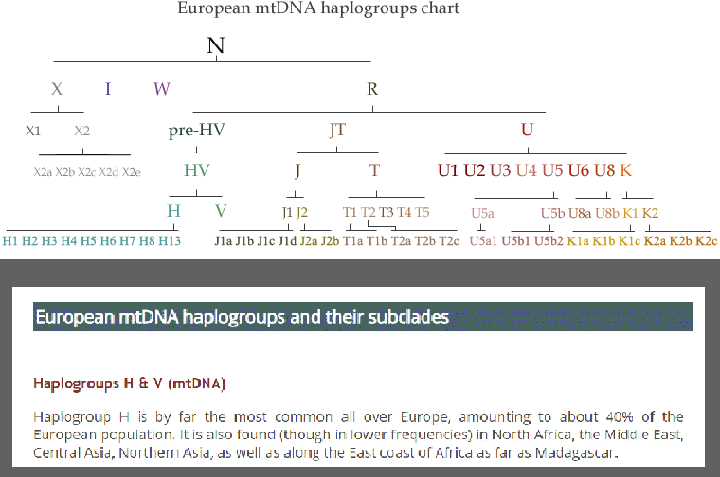

The female H haplogroup from Eupedia.com on haplogroups
Men have a Y chromosome, which makes them male, which has been passed from father to son, to his son, to his son, and so forth from from time immemorial. We all have mitochondrial DNA (mtDNA ) which is passed from a mother to all her children unchanged. Thus your mtDNA is from your mother’s mother ‘s mother and so on. Both of those parts of DNA inheritance can be traced back to the dawn of humanity. That is unlike the other chromosomes which mix the inheritance from each parent such that after several generations there may be little or no trace of our deeper ancestors. Most of us have no verifiable autosomal DNA from before our 5th grandparents.
Those of you who have family legends about descent from an Indian princess might be able to prove the connection using mtDNA if there is a direct female line to that ancestor, since there are specific haplogroups for Native Americans (click here for the wikipedia article on that).
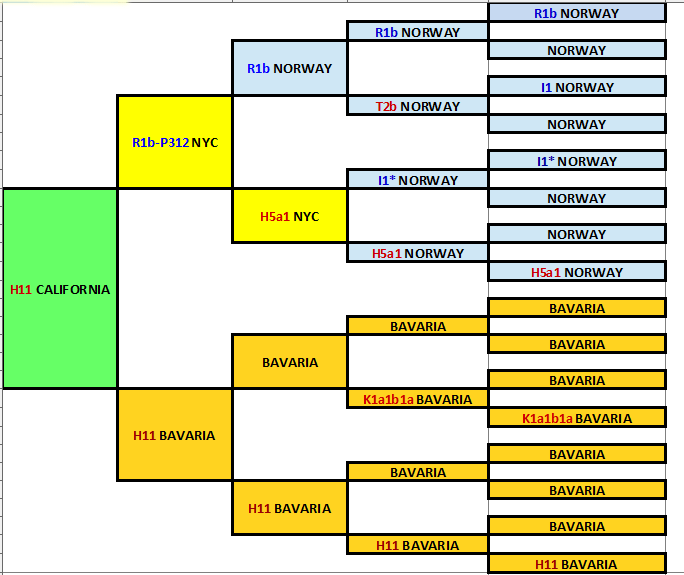
My Ancestral Haplogroups displayed in Paul Hawthorne’s colorful genealogy chart
One thing that I like to do is figure out the haplogroups of my recent ancestors by testing cousins in the needed line of descent. I made a chart of the ones I know using Paul Hawthorne’s colorful genealogy chart (click here for more about that) with the haplogroups added. As you can see, I have many more lines to chase down. Sadly my Thannhauser Bavarian Jewish line daughtered out, so I am trying to find a male descendant of the one who moved to Albany NY in the mid 1800s.
So how do you find your haplogroup from your DNA test? Well if you tested at 23andme or Living DNA then you will be provided with your high level haplogroup. However if you want to drill down the branches, then test your Y and/or your mtDNA at Family Tree DNA (summer sale until end of August). Ancestry tests enough SNPs to get a high level haplogroup by using other tools on your raw data. My Why Y blog post explains how to use the Morley tool but there is also a tool to find Y haplogroups from Borland Genetics. I have been trying to convince Kevin Borland to write one for mtDNA since the James Lick mthap tool will not currently take ancestry data.